 coli K12 with various input ratios and average cell concentrations (C avg) using MEGAs. High-throughput digital multiplex detection of E. For all tests of droplet generation, the mock PCR mix containing ~100 beads per uL was used. coli K12 at 0.2 cells per droplet (cpd) in 2.5 nL droplets. Right: a representative flow cytometric histogram of beads carrying the FAM labeled PCR product from E. Middle: size distribution of uniform ~2 nL droplets collected from sixteen nozzles in a device. Left: image of droplet production at the T-shaped nozzles with a total throughput of 2.4 × 10 6 droplets per hour. Right: a representative flow cytometric analysis of 624 bp amplicons on beads amplified from pUC18 template (1 copy/droplet) in ~3 nL droplets generated from four nozzles in a device. Middle: size distribution of uniform droplets collected from eight nozzles in a device. Left: photo of the cross-shaped nozzles each generating ~4 nL droplets at 5.6 Hz (6.4 × 10 5 droplets per hour). This result demonstrates that multiplex SCGA is a promising tool for high-throughput quantitative digital analysis of genetic variation in complex populations.Ĭharacterization of droplet generation and PCR amplification using MEGAs. coli O157 cells in a high background of normal K12 cells, with a detection limit on the order of 1/10(5). 
Using Escherichia coli bacterial cells as a model, we show that this technique enables digital detection of pathogenic E. 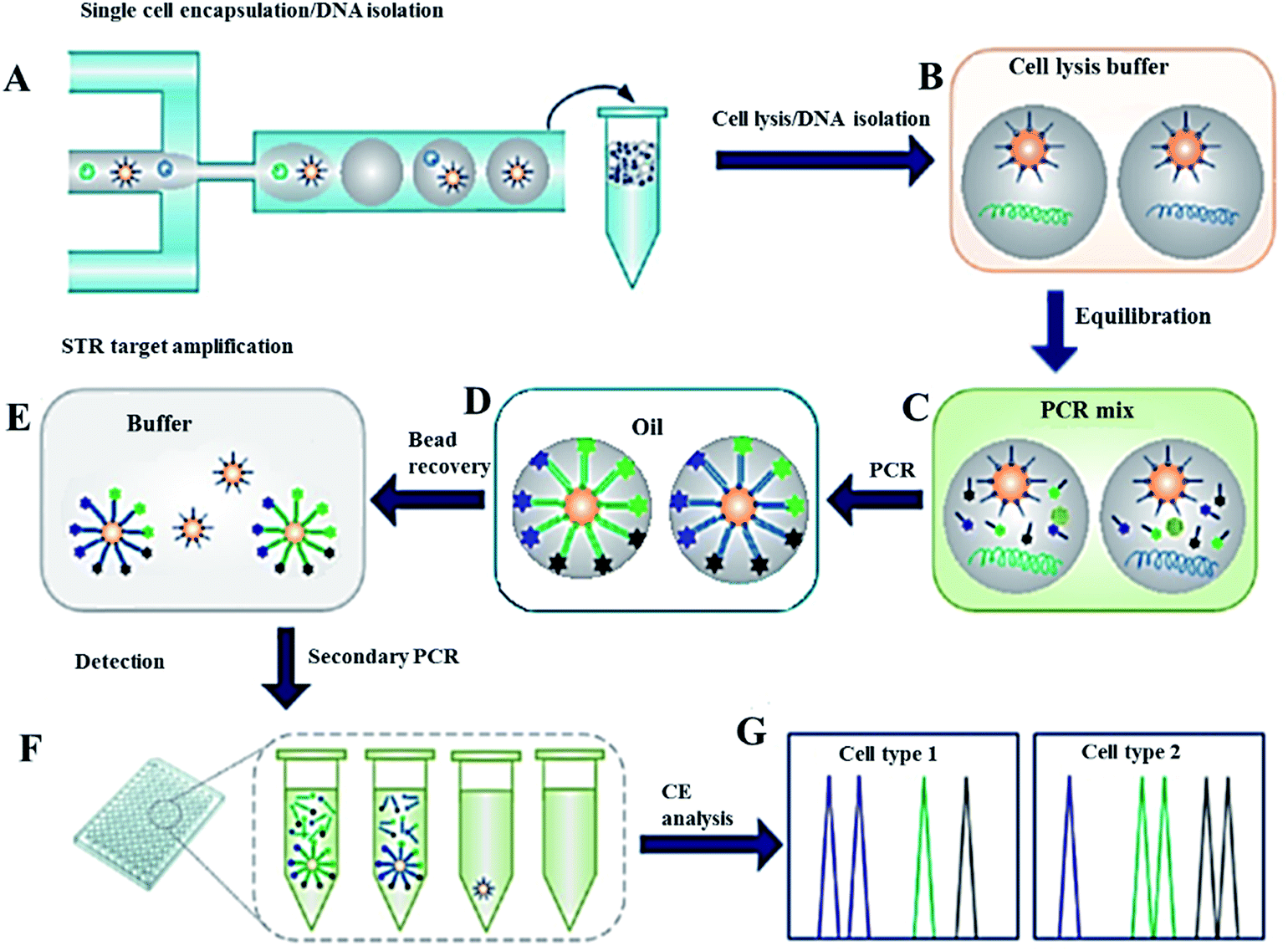
Following PCR, the droplets are lysed and the beads are pooled and rapidly analyzed by multicolor flow cytometry. In this method, microbeads functionalized with multiple forward primers targeting specific genes from different cell types are used for solid-phase PCR in droplets. A multiplex single cell PCR method is developed to detect and quantify both wild type and mutant/pathogenic cells. Hybrid glass-polydimethylsiloxane diaphragm micropumps integrated into the MEGA chips afford uniform droplet formation, controlled generation frequency, and effective transportation and encapsulation of primer functionalized microbeads and cells. Microfabricated emulsion generator array (MEGA) devices containing 4, 32, and 96 channels are developed to confer a flexible capability of generating up to 3.4 x 10(6) nanoliter-volume droplets per hour. Here we describe a high-performance single cell genetic analysis (SCGA) technique that combines high-throughput microfluidic emulsion generation with single cell multiplex polymerase chain reaction (PCR). High-throughput genetic and phenotypic analysis at the single cell level is critical to advance our understanding of the molecular mechanisms underlying cellular function and dysfunction. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
